Keywords
biomarker, image processing, cell segmentation, tilebased, processing, fluorescence in situ hybridization (FISH), region, merging.
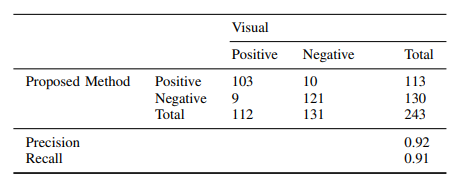
Quantitative output variables
HER2/neu gene status has become indispensable in pathologic scoring of invasive ductal breast carcinoma. Although FISH is an expensive and time consuming process, it guarantees the result of the case. If the result is positive, Trastuzumab therapy is given to the patient. False treatments with Trastuzumab lead to loss of money and time. It is also essential to be sure that the patient morality is secured by avoiding false treatments. Our proposed algorithm has three advantages. Basically, it includes whole slide processing that lets user control all parts of the slide. It does not require color phase adjustment such as adjustment of brightness and contrast, and it is also very fast and robust due to fact that we divided each slide into tiles, which reduces the memory need.
References
[1] V. Ozmen, "Breast cancer in turkey: clinical and histopathological ¨ characteristics (analysis of 13.240 patients)," The journal of breast health, vol. 10, no. 2, p. 98, 2014.
[2] N. Iqbal and N. Iqbal, "Human epidermal growth factor receptor 2 (her2) in cancers: overexpression and therapeutic implications," Molecular biology international, vol. 2014, 2014.
[3] B. Lerner, "Bayesian fluorescence in situ hybridisation signal classification," Artificial Intelligence in Medicine, vol. 30, no. 3, pp. 301–316, 2004.
[4] F. Raimondo, M. A. Gavrielides, G. Karayannopoulou, K. Lyroudia, I. Pitas, and I. Kostopoulos, "Automated evaluation of her-2/neu status in breast tissue from fluorescent in situ hybridization images," IEEE Transactions on Image Processing, vol. 14, no. 9, pp. 1288–1299, 2005.
[5] H. Masmoudi, S. M. Hewitt, N. Petrick, K. J. Myers, and M. A. Gavrielides, "Automated quantitative assessment of her-2/neu immunohistochemical expression in breast cancer," IEEE transactions on medical imaging, vol. 28, no. 6, pp. 916–925, 2009.
[6] E. M. van der Logt, D. A. Kuperus, J. W. van Setten, M. C. van den Heuvel, J. E. Boers, E. Schuuring, and R. E. Kibbelaar, "Fully automated fluorescent in situ hybridization (fish) staining and digital analysis of her2 in breast cancer: a validation study," PloS one, vol. 10, no. 4, p. e0123201, 2015.
[7] S.-C. Huang, F.-C. Cheng, and Y.-S. Chiu, "Efficient contrast enhancement using adaptive gamma correction with weighting distribution," IEEE Transactions on Image Processing, vol. 22, no. 8, pp. 1032–1041, 2013.
[8] A. M. Reza, "Realization of the contrast limited adaptive histogram equalization (clahe) for real-time image enhancement," Journal of VLSI signal processing systems for signal, image and video technology, vol. 38, no. 9, pp. 35–44, 2004.
[9] S. Razavi, G. Hatipoglu, and H. Yalc¸ın, "Automatically diagnosing her2 ˘ amplification status for breast cancer patients using large fish images," in Signal Processing and Communications Applications Conference (SIU), 2017 25th, no. 10, pp. 1–4, IEEE, 2017.
[10] S. Fontenete, D. Carvalho, A. Lourenc¸o, N. Guimaraes, P. Madureira, ˜ C. Figueiredo, and N. F. Azevedo, "Fishji: New imagej macros for the quantification of fluorescence in epifluorescence images," Biochemical engineering journal, vol. 112, no. 11, pp. 61–69, 2016.